|
If the positive gate contains less than 2%, try setting a parent (preceding) gate to ensure that your positive gate meets or exceeds 2%.Ĩ) You can also create your own gates in the graph window and use the text assignment windows, within the compensation editor, under the negative and positive columns in the top portion of the compensation interface to assign these newly created gates.ĩ) Please name the compensation matrix. Also, there are rules regarding the positive population – it must contain more than 100 events and be 2% or greater of the parent population. However, if there is a noticeable problem, manually alter the gate position. In most cases, these gates settings will be optimized. *The compensation algorithm determines gate locations empirically based on fluorescence. When you double click, the graph window will open and all you to adjust the location of the gates. 3 of the 7 gated and compensated parameters are shown below, highlighted by the red box.ħ) Double click on a graph to edit a gate location. You can also use this menu to add additional parameter options (if they were missed by the compensation editor) or to remove parameter options that may be complicating the interface.Ħ) To adjust any gates, the bottom portion of the window provides a list of each parameter, with graphs of gates created by FlowJo for scatter and positive/negative populations for each single stain control. As you can see from the picture below, the CD3-7AAD parameter and APC 647 require adjustment.ĥ) You can use the assignment text at the top of the window to clear sample assignments if they are incorrect. Green indicates a good match, yellow is less certain, and red indicates a poor or non-existent control file is associated the compensation parameter. A colored dot will appear next to the file name indicating the likelihood that FlowJo found the correct compensation file. 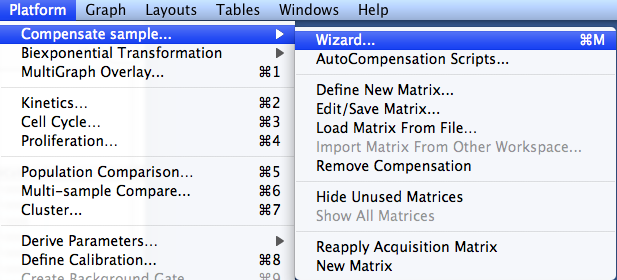
An overview of this window is here.Ĥ) FlowJo will attempt to assign the correct samples to each parameter. Otherwise, the compensation interface will not process the controls correctly.ġ) To initiate creating a new compensation matrix in FlowJo, select the compensation group in the workspace and go to the Workspace ribbon.Ģ) Click the compensation icon in the Cytometry band of the Tools tab.ģ) The compensation interface will launch. Only put the controls necessary for creating that compensation matrix in this compensation matrix. One compensation group will yield one compensation matrix from the controls provided to that group. If you require multiple new compensation matrices, simply create a new group and assign it a role of ‘compensation’. 
There is a single compensation matrix in the workspace by default and it will allow you to create one new compensation matrix in FlowJo. Compensation controls (single stain samples) must now belong to a compensation group in order to create a compensation matrix with these controls. 
Question: Did the Diva Software or our Device mess up the FCS file for some reason or is a minor inconsistency in the fcs file treated very strictly by read.Chapter 2 – Creating a New Compensation Matrix in FlowJo CHAPTER 2 – Creating a FlowJo compensation matrix from single stain controlsīefore compensating data in FlowJo, be sure your parameters which require compensation are scaled correctly.Ĭompensation has been reworked for version 10. So, somehow FlowJo is able to fix the file but of course I would like to understand the underlying problem. When I import the File to FlowJo and then export it again, read.FCS(x) works. Interestingly, FlowJo does not show the compensation matrix as usual but the Spill-Matrix is in the FCS file's keywords or parameters. In comparison to other files that I have and that can be read by read.FCS(x) this file was subject to manual compensation (not the compensation wizard) with the FACS Diva Software on a BD device. $SPILLOVER size discrepancy could not be resolved! $SPILLOVER keyword value is of improper size for number of spillover channels! When calling read.FCS(x) an error occurs:
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
